Some Mammograms contain the medical clips / marker / insert proper terminology here - and these cause issues when attempting to Congeal different scans, as these clips are recognised, and they try to align with each other also. This may detract from the alignment of the breast tissue.
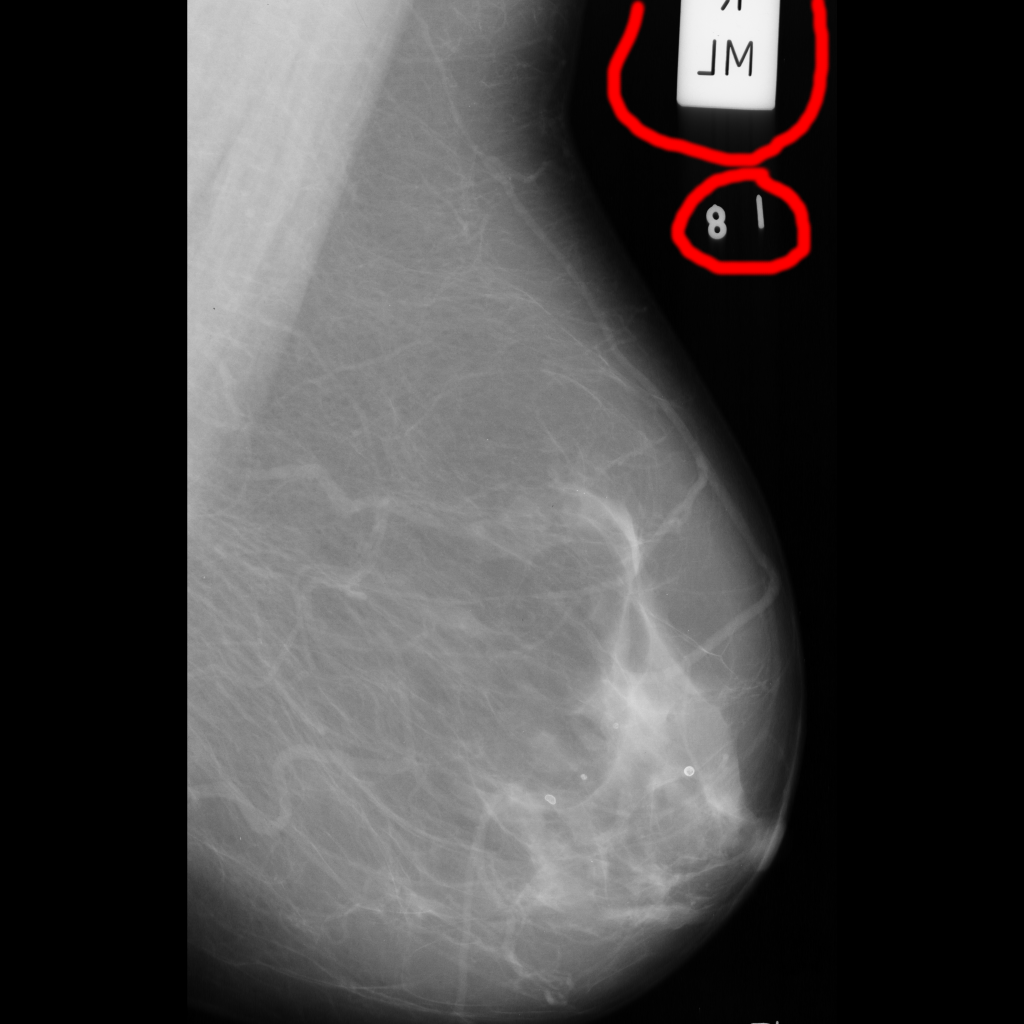 Not sure what the numbers are? Part of the marker maybe?
Not sure what the numbers are? Part of the marker maybe?
So, to reduce this issue I have two options:
- Ask the user to use only scans without markers
- Limiting as I'm already asking them to use the same orientation of scan
- Remove these labels using some image processing wizardry
- Very lack in expertise in this area - but worth a shot!
So let's look through some of the methods I have investigated, and how well their faired.
Morphological Operations - looking for squares
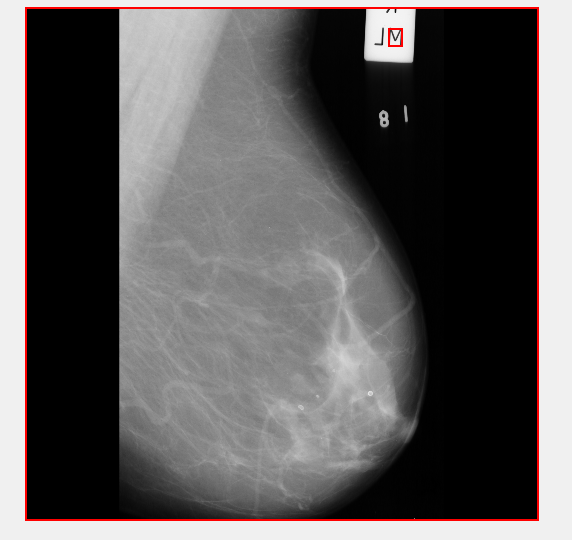
As is pretty evident here - this isn't going to work. The Marker itself isn't square, however the image is! So if I remove squares from this image - well goodbye image!
bwareaopen
This is a nice little function in MATLAB which can remove small objects from images - such as noise after conversion to black and white.
Read in the image, and remove all objects fewer than 'x' pixels.
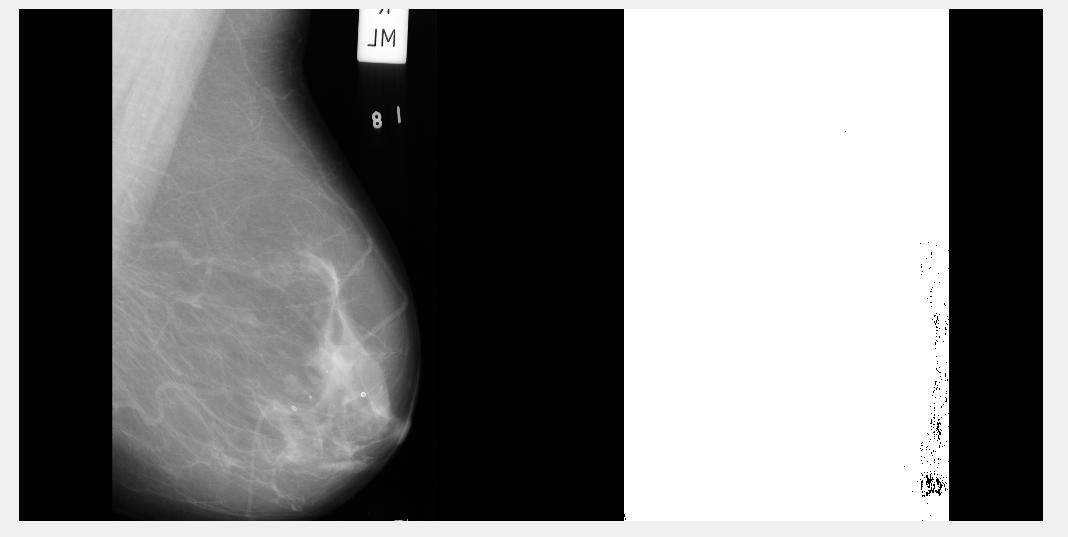
Left image is the original scan, right image is after bwareaopen is run.
Not even sure what's happened here - no amount of tweaking 'x' value made a difference. I can only presume this is because breast tissue is a variety of grey-scale values?
Not sure... If anyone knows why, let me know!
regionprops
This MATLAB function apparently 'Measure properties of image regions' - I wasn't really sure what I'd get out of this, but it was recommended right next to bwareaopen for removing objects from images.
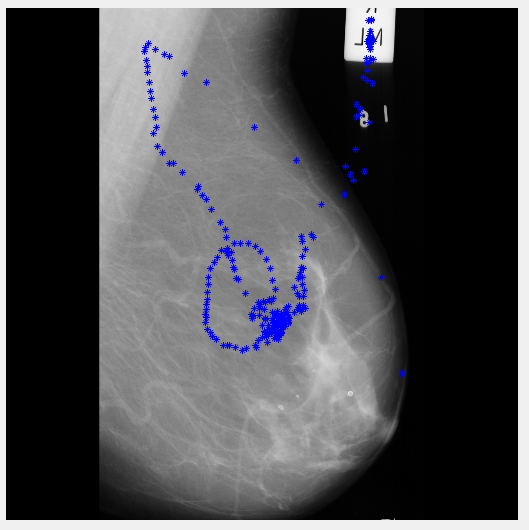
After running the algorithm, I'm still not sure what it's doing. Not an edge detector obviously, seems a bit sporadic in places - but it does go up to the label!
Maybe if I can figure out what this is actually showing I might be able to use it...
Shape recognition demo
I then stumbled across this nice little demo that Ahmed Samieh put online in 2007.
My hope with this one was to detect the straight edges of the label - from this I could look to removing it. The demo took a little tweaking as my input image was already grey-scale, not RGB as his was.
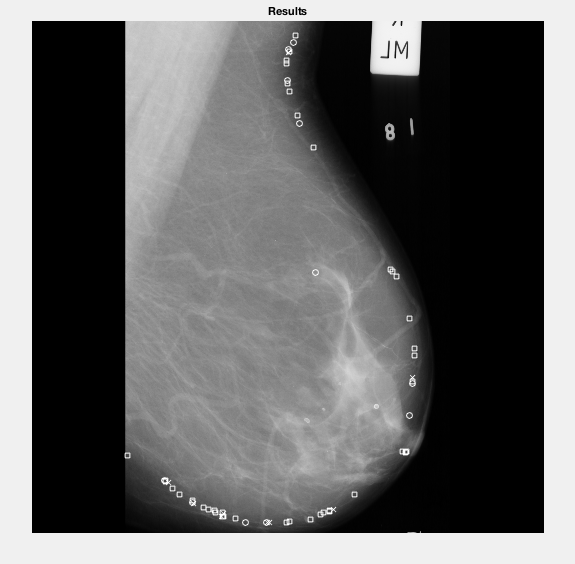
As you can see it actually didn't even pick up the label, however it did pick up the breast tissue.
It's not quite accurate enough in places, trims pieces off the edge and has 1 random point in the breast tissue - however had it been more accurate I possibly could've used this to then remove the background from the image.
Removing white areas?
So after some in-depth googling, I stumbled across a thread where someone wanted to remove the glare from a jewellery photo.
Someone called 'Image Analyst' (who is this person? They crop up everywhere!) replied with a demo on removing white objects from an RGB photo.
Again, through some tweaking of this I came across something very interesting.
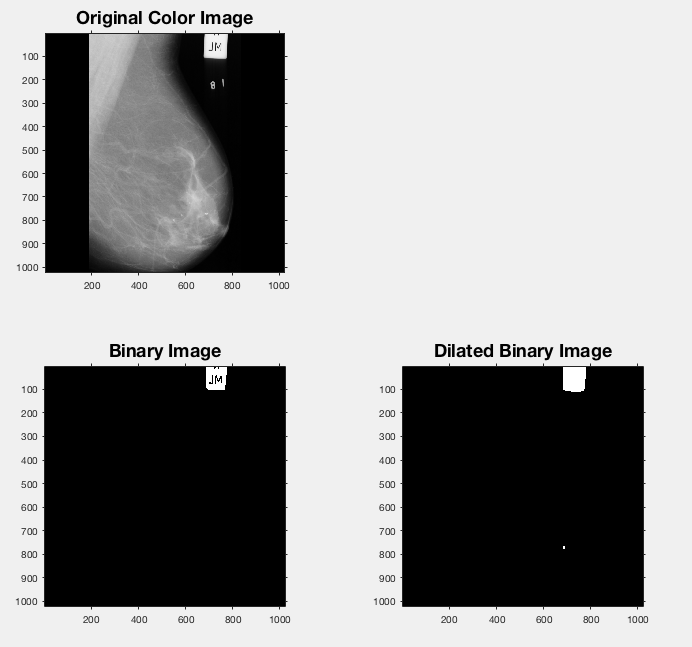
The 'Binary Image' they create in their demo seemingly picks out the label!
Turns out creating this binary image was actually just taking the original image (or a grey-scale version of it if it's RGB) and looking for values greater than 225 (I tweaked this a bit to see if I could get a better value in my own example - settled on 220).
Image Subtraction
So having this label picked out is great, but now I want to remove it from the original image. Guess what? MATLAB has something for that too! A neat little function called imsubtract.
So I subtract the binary image (after converting it back to uint8 from logical) and here's the output:
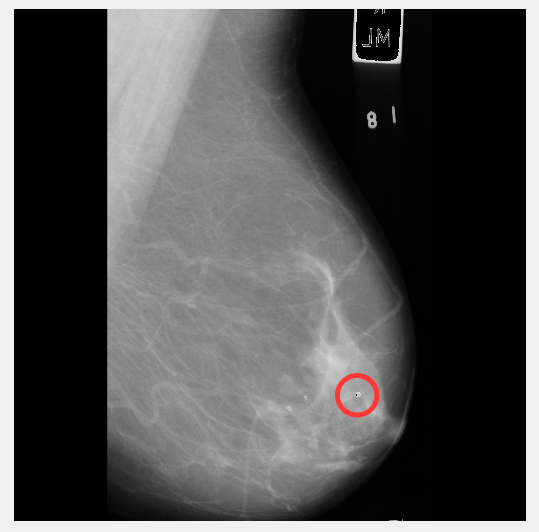
Not bad apart from that small bit circled.
What about if we lower the grey-level we're removing? So all items over 200 are removed - maybe the label edge will be removed?
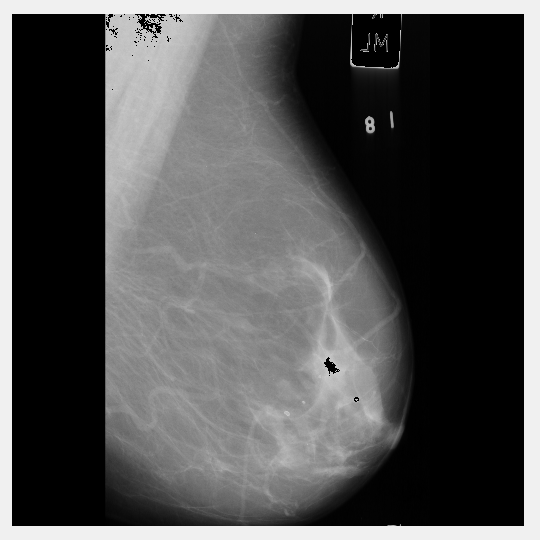
No such luck - it starts to remove vital breast tissue from the scan. Back to 220 grey-level removal it is.
So what about on a different scan?
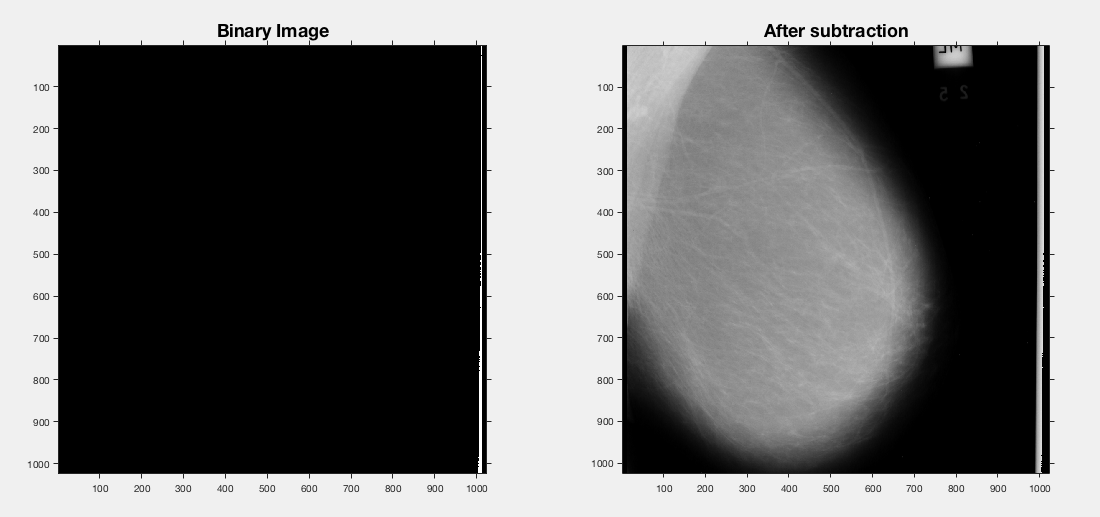
No such luck - items above 220 don't pick up anything in this scan.
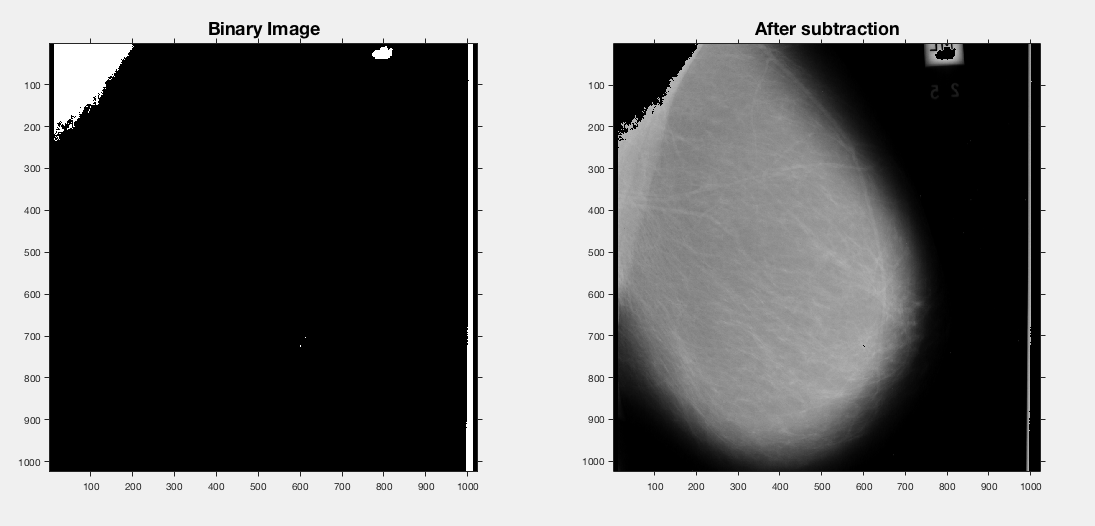

Lower the grey-level removal to 180 and the label is starting to be removed - however we're also starting to lose breast tissue also.
Whilst this is the best approach I have found as of yet - it isn't perfect. It's down to a case per case basis, meaning the User would have to adjust the grey-level values manually and experiment with the best fit before saving each individual image back into the folder for combining into one large image.
The rest of today (Monday 28th March) will be spent investigating into this and I will update the post if I find a solution!
 Post-edit
Once again stumbling across a nice little demo by Image Analyst (who are you??) I think I've found one solution to removing the labels.
Using the tool imfreehand in MATLAB, over the input image, you can select a 'Mask' which you can remove from the image.
The way in which it removes is to set the mask to 0 (absolute black) - so my only concern is that the background of the scans mightn't be absolute black, and it will still be anomaly in the image. Only experimentation will prove that I guess?
Here's some screenshots of Image Analyst's demo (slightly tweaked by me to fix a bug in maximising the figure) :
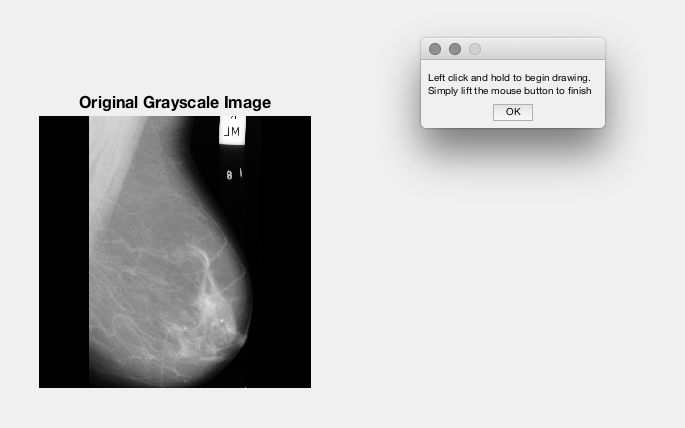

So what I could do is offer an option to the User to open a new GUI over the top of the existing if they wish to remove any markers. They could load in the image they wish to edit, edit it, and save it back out to the same location - overwriting the image would be the only option given the current situation.
Not a perfect solution - and indeed I mightn't include it in my final project - but it's possibly better than congealing markers.